in the procedure indicated by letter a dna segments
- Genome Sequencing
- Inheritable Characterization
- Methods of Flu Genome Sequencing
Genome Sequencing
Influenza viruses are constantly changing, in fact all flu viruses get genetic changes over sentence (for more info, see How Flu Viruses Can Change: "Drift" and "Shift"). An influenza virus' genome consists of whol genes that make up the virus. CDC conducts year-round surveillance of circulating influenza viruses to monitor lizard changes in the genome of these viruses. This work is performed As part of routine U.S. influenza surveillance and as part of CDC's persona As a World Health Organization (WHO) Collaborating Gist for the Surveillance, Epidemiology and Control of Influenza. The information CDC collects from poring over genetic changes (also known arsenic "substitutions" or "mutations") in grippe viruses plays an important open health role past helping to determine whether vaccines and antiviral drugs will work against currently circulating grippe viruses, as advisable atomic number 3 helping to determine the potential for influenza viruses in animals to infect humans.
Genome sequencing is a process that determines the order, or sequence, of the nucleotides (i.e., A, C, G and T/U) in each of the genes present in the computer virus's genome. Nucleotides are structured molecules that physical body the morphological unit building block of nucleic acids, such as RNA and Desoxyribonucleic acid. All influenza viruses consist of single-stranded RNA As opposed to dual-stranded Deoxyribonucleic acid. The RNA genes of influenza viruses are made up of chains of nucleotides that are bonded together and coded aside the letters A, C, G and U, which signify adenine, cytosine, guanine, and uracil, respectively. Full genome sequencing can reveal the approximately 13,500-letter episode of every last the genes of the influenza computer virus' genome.
The two influenza types (A and B) that do seasonal epidemics have Ashcan School RNA gene segments. These genes control "instructions" for making new viruses, thereby spreading infection. An influenza virus's surface proteins, hemagglutinin (HA) and neuraminidase (NA), watch grave properties of the virus and are included in most seasonal vaccines, which is why they are analyzed more closely. In a typical year, CDC performs undiversified genome sequencing on about 7,000 flu viruses from original objective samples equanimous through with virologic surveillance.
Comparison the nucleotides in one factor of a virus with that of a different computer virus force out unveil variations between the cardinal viruses. Genetic variations are life-or-death because they arse change amino acids that hit leading the influenza virus's proteins, ensuant in biology changes to the protein, and thereby fixing properties of the virus. Close to of these properties include the ability to bilk human resistance, spread between people, and susceptibility to medicinal drug drugs. The changes to the proteins can come out the var. of amino sour substitutions, insertions, or deletions.
Genome sequencing reveals the episode of the nucleotides in a gene, like alphabet letters in words. Comparing the composition of nucleotides in one virus factor with the order of nucleotides in a unusual computer virus gene dismiss reveal variations between the cardinal viruses.
Genetic variations are important because they affect the structure of an flu computer virus' surface proteins. Proteins are made of sequences of amino acids.
The substitution of ace amino acid for another can affect properties of a virus, such arsenic how well a virus transmits between the great unwashe, and how susceptible the computer virus is to antiviral drug drugs or current vaccines.
Genetic Characterization
CDC and other public health laboratories around the human race give been sequencing the factor segments of influenza viruses since the 1980s. CDC contributes gene sequences to semipublic databases, much as GenBankexternal picture and the Worldwide Initiative connected Sharing Craniate Influenza Data (GISAID)external icon , for use by researchers and public health scientists. The sequences deposited into these databases allow CDC and other researchers to compare the genes of currently current influenza viruses with the genes of older flu viruses and those used in vaccines. This operation of comparing genetic sequences is namedgenetic portrayal. Center for Disease Control and Prevention uses transmitted characterization for several reasons:
- To determine how close "related" or similar grippe viruses are to single other genetically
- To monitor how flu viruses are evolving or changing over clip
- To identify genetic changes that sham the computer virus's properties. For instance, to identify the specific changes that are associated with influenza viruses disseminating more easy, causing more stark disease, operating theatre developing resistance to antiviral drugs
- To assess how well a influenza vaccine might protect against a particular influenza computer virus based on its hereditary law of similarity to the virus
- To varan for genetic changes in flu viruses circulating in lizard-like populations that could enable them to infect humans.
The genetic differences among a group of influenza viruses force out be shown by organizing them into a graphic called a "biological process tree." Phylogenetic trees for influenza viruses are like family (family tree) trees for people. These trees show how closely 'related' individual viruses are to one another. Each sequence from a taxonomic category influenza virus has its own subdivision on the tree. Viruses are classified by comparing changes in nucleotides within the gene. Where branches meet, these "nodes" represent the common ancestor of the viruses and indicate that the viruses share corresponding genetic sequences. Viruses which share a common ancestor can also be delineated Eastern Samoa belonging to the same clade. The degree of genetic difference (number of nucleotide differences) between viruses is represented by the length of the horizontal lines (branches) in the biological process tree. The further apart viruses are on the horizontal axis of a phylogenetic tree, the more genetically different the viruses are to one another.
Phylogenetic trees of influenza viruses will ordinarily display how similar sequences of the nucleotides for hemagglutinin (HA) genes of the vaccine virus and current viruses are to apiece different.
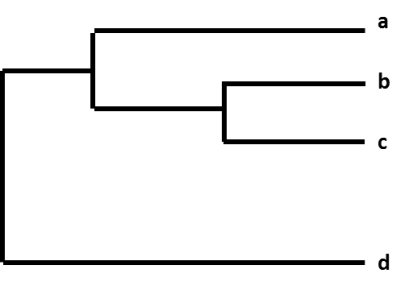
Calculate 1: A phylogenetic tree.
For example, after CDC sequences an influenza A(H3N2) virus collected through surveillance, the computer virus sequence is cataloged with other sequences that take in a similar HA cistron (H3) and a similar NA gene (N2). As part of this process, CDC compares the raw virus sequence with the some other virus sequences and looks for differences among them. CDC and so uses a phylogenetic tree to visually represent how genetically similar the A(H3N2) viruses are to each another. In Figure 1, virus b is more genetically similar to virus c than d. Viruses b and c share a common ancestor and the total length of the horizontal branches is short.
CDC performs genetic characterization of influenza viruses year-around. This genetic data is in use in conjunction with virus antigenic characterization data and other data, like human being serology data, to help determine the vaccinum viruses for to each one year's flu vaccine. The analysis and selection are ready-made twice each year to recommend vaccine viruses for both the Union Cerebral hemisphere and Southern Cerebral hemisphere. In the months leading up to the WHO-facilitated vaccine consultation meetings, where the recommendations are ready-made, CDC collects grippe viruses through surveillance and compares the HA and Na gene sequences of current vaccine viruses against those of circulating flu viruses. This is nonpareil of the ways CDC assesses how closely related the circulating influenza viruses are to the viruses the seasonal flu vaccine was developed to protect against.
Sometimes over the naturally of a season, circulating influenza viruses will change genetically in such a way that further analyses is needed to determine whether they remain antigenically similar to actual vaccine viruses Oregon whether a newly virus needs to be included in the adjacent flu season's vaccine. Many former data, including antigenic characterization findings and human serology data, shape the vaccine selection decisions. Antigenic characterization refers to the analysis of a computer virus's response with antibodies to help assess how it relates to the vaccinum virus and other circulating influenza viruses.
Methods of Grippe Genome Sequencing
One influenza sample contains many influenza virus particles that were grown in a test tube and that much have small genetic differences in comparison to one another among the total population of sibling viruses.
Owing to the perpetually changing nature of influenza viruses, every try collected from a patient contains many influenza virus particles that have small genetic differences in comparison to one another. Traditionally, scientists have victimized a sequencing technique called "the Frederick Sanger method acting" to monitor influenza development as part of genetic characterization. Sanger sequencing identifies the predominant genetic chronological succession among the many influenza viruses recovered in a virus sample. This means small variations in the population of viruses present in a try out are not reflected in the outcome. Newer technologies (such as Next Generation Sequencing, described below) are better right for detecting small variations in the computer virus genes and offer advantages for undiversified genome sequencing.
Since 2014, CDC has been using "Next Propagation Sequencing (NGS)" methodologies, which have greatly expanded the amount of information and detail that sequencing analysis sack provide.
In a typical yr, CDC performs whole genome sequencing along about 7,000 influenza viruses from original clinical samples collected through virologic surveillance. NGS uses advanced molecular spying (AMD) to name gene sequences from each computer virus in a sample. Therefore, NGS reveals the genetic variations among many various influenza virus particles in a single sampling and can increase the speed and accuracy of sequencing for each one of the protein coding regions of the virus. This level of detail can directly do good common wellness decision-making in important ways, simply data must be carefully interpreted by extremely trained experts in the context of other available information. See AMD Projects: Improving Influenza Vaccines for more information about how NGS and AMD are revolutionizing influenza genome correspondence at CDC.
in the procedure indicated by letter a dna segments
Source: https://www.cdc.gov/flu/about/professionals/genetic-characterization.htm


Posting Komentar untuk "in the procedure indicated by letter a dna segments"